When the real thing—in this case, the human brain—is rightfully out of the reach of experimental manipulation, what’s the next best thing for understanding how the brain works in health, falters in disease, and could be rescued with treatment? The longtime answer has been animal models, and often that’s still true. But pioneering neuroscientists and engineers are advancing another way: simulation.
Among this vanguard are Picower Professors Li-Huei Tsai and Earl K. Miller, who have each forged collaborations to produce novel simulations of the brain. Each has recently published papers about these inventions, though their teams’ ongoing work means they’ve already surpassed those milestones and developed the technologies further.
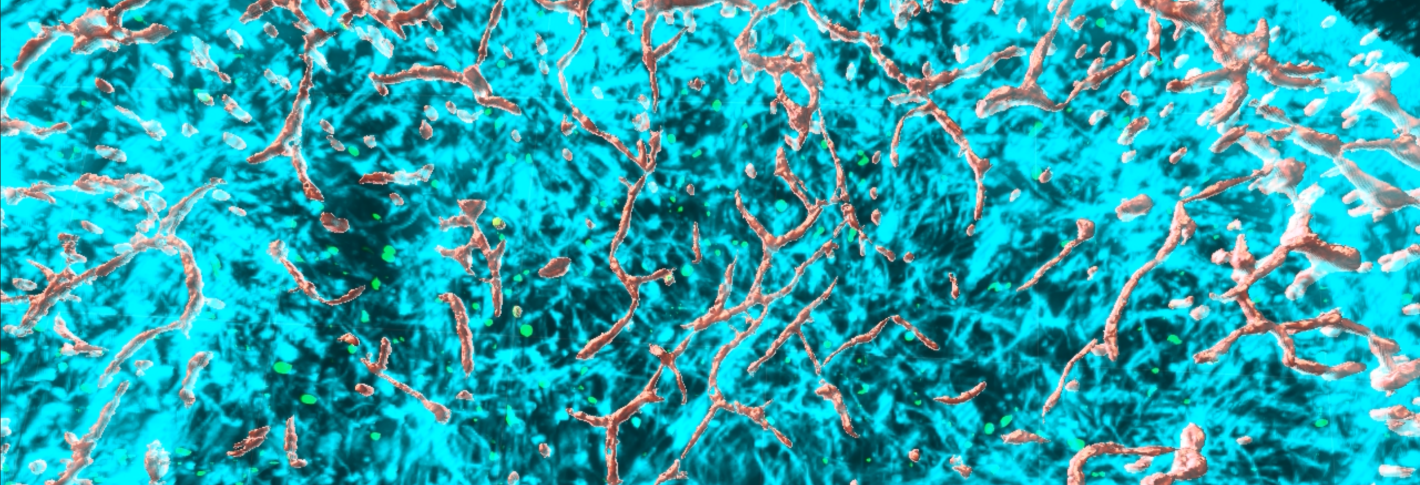
In Tsai’s case, the innovation is the “miBrain,” the first 3D human brain tissue culture to include all six major brain cell types. Demonstrating that the unprecedentedly diverse mini-tissue is an advanced platform for discovery, Tsai and colleagues used it to produce a new revelation about how cellular interactions drive Alzheimer’s disease. In Miller’s case, the mental mimic is a biologically-based computational model of brain circuitry and dynamics called “Neuroblox.” His team showed it can replicate learning behavior and revealed neural activity in its simulated circuits that had gone unnoticed in real brain data.
Each simulation enables new paradigms of experimental manipulations to yield fundamental discoveries. They also provide advanced high-volume platforms for testing interventions and treatments to improve brain health.
“The idea is to make a platform for biomimetic modeling of the brain so you can do things like drug development and drug testing earlier in the process, providing a more efficient way of developing neural therapies,” said Miller, one of the six founders of the Neuroblox startup company led by CEO Lilianne R. Mujica-Parodi.
Like Neuroblox, miBrain offers new research options, Tsai said.
“The miBrain can develop human-specific pathological features without ectopically overexpressing a particular human disease gene, as one must do with animal models of human diseases,” Tsai said. “Also, it is possible to create personalized miBrains using skin cells donated by patients. Finally, the miBrain culture is very amenable to drug treatment. Therefore, the miBrain can be used as a parallel system to animal models.”
A culture of ingenuity
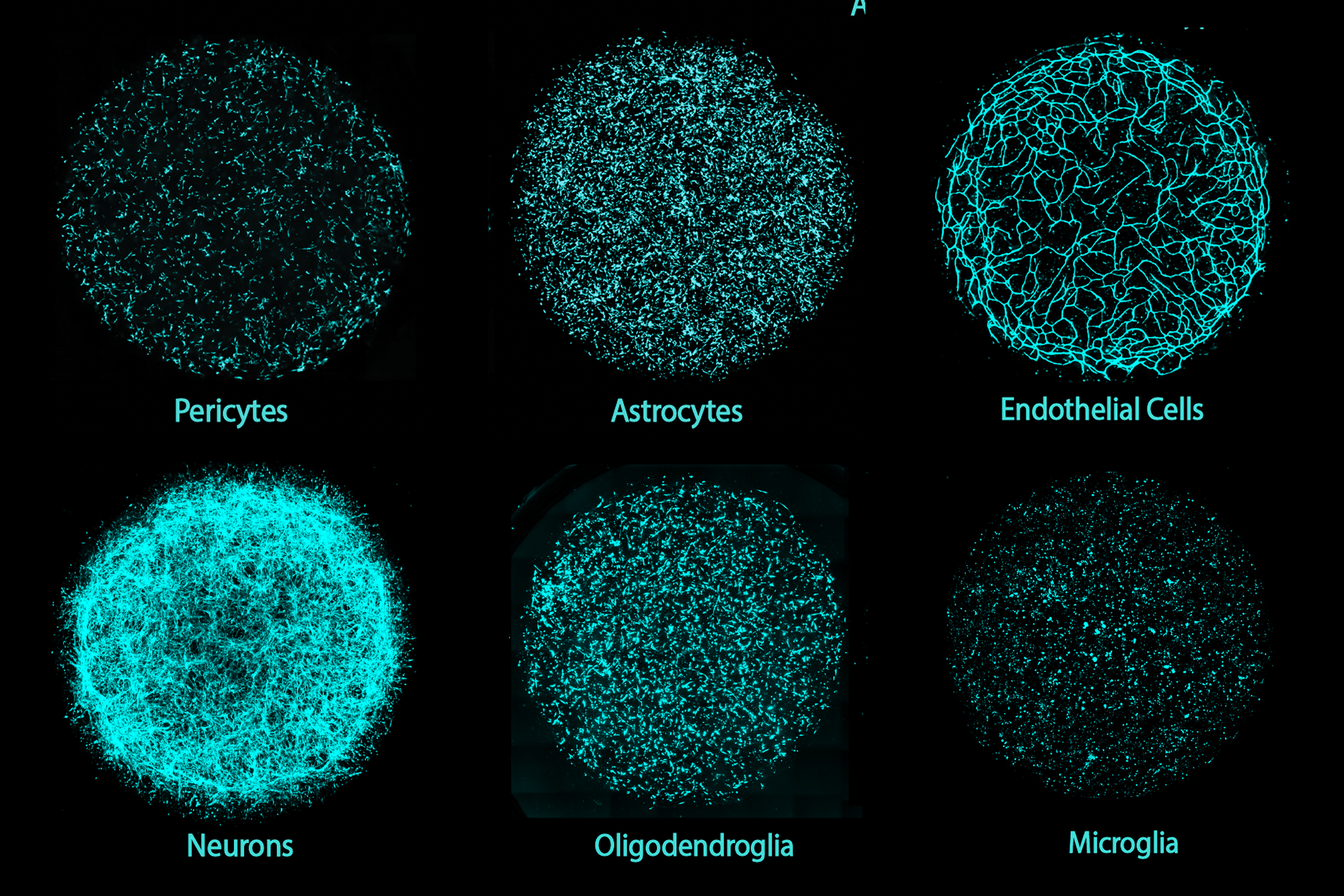
For years, Tsai’s lab has been using 3D human cell cultures (as have Picower colleagues Kwanghun Chung and Mriganka Sur). But keenly aware of how complex the brain, and the Alzheimer’s pathology that afflicts it, can be, Tsai sought to integrate all six major cell types: neurons, microglia, oligodendrocytes, astrocytes, pericytes and endothelial cells.
The work began in 2019 with a grant that Tsai and former postdoc Joel Blanchard earned from the National Institutes of Health. They teamed up with the chemical engineering and bioengineering lab of MIT Institute Professor Robert Langer. In an October 2025 paper in the Proceedings of the National Academy of Sciences, led by former postdocs Alice Stanton and Adele Bubnys, they published their landmark result: the multicellular integrated brain, or miBrain.
All cells in a miBrain derive from skin cells donated by a volunteer. Those cells are converted back into stem cells that are then transformed into the different brain cell types. Then they are combined into the full culture within an ingenious hydrogel. As such, each miBrain is a personalized model brain that carries the genome of the donor and self-assembles into functioning units, including blood vessels, immune defenses, myelinated axons, and neural circuits.
Meanwhile, the lab can grow many copies of someone’s miBrain and alter any of the genes in the cells. They can also expose the miBrains to various drugs or other agents to test their effects, making them a scalable platform for personalized treatment screening.
In the PNAS paper, the team used miBrain to study how the APOE4 variant of the APOE gene, the biggest genetic risk factor for Alzheimer’s, produces pathology such tau proteins. The role of astrocyte cells is an open question that couldn’t be fully resolved just by looking at post-mortem brains or by working with a culture of astrocytes alone.
Indeed, when the team cultured just APOE4 astrocytes, they found that the cells did not exhibit the same pathology-promoting immune response in lab tests that they produced in miBrains where they interacted with their fellow brain cells. Yet, pathology occurred in miBrains where the astrocytes harbored APOE4 but all the other cell types harbored the benign APOE3 version. That suggested that APOE4 compels astrocytes to promote pathology, but only in conjunction with other cells.
Data from previous studies hinted that the APOE4 astrocytes’ partners in pathology might be microglia cells, so the researchers left microglia out of some miBrains. In those cases, APOE4 astrocytes did not promote pathology. And when they exposed microglia-less miBrains to doses of media from a microglia-astrocyte co-culture, they saw tau increase, but not when they seeded such miBrains with media from just microglia or astrocyte cultures alone. The modularity of miBrains allowed them to show that APOE4 astrocytes depend on interaction with microglia to promote Alzheimer’s pathology.
The team has extended miBrains even further. In a preprint, the team describes clever modifications that have enabled the cultures to grow circulatory networks. By hooking them up to microfluidic systems, the scientists can perfuse the circulatory system to test therapeutic agents, including how they might fare at the formidable “blood-brain barrier” (BBB) that strictly filters what goes into or out of the brain from its blood vessels.
“The perfusable BBB allows us to introduce protein factors, peptides, chemicals and drugs into the miBrain,” Tsai said. “It also permits the potential evaluation of pharmacokinetics and pharmacodynamics of drugs such as how efficiently they can cross the BBB.”
Brainy blocks
Neuroblox, too, is a fusion of neuroscience and engineering. About five years ago, Mujica-Parodi realized that neuroscience studies and therapies could be accelerated if the field had the computer simulation tools that benefit computer chip or aerospace designers. Computational models free experimenters from logistical constraints of working with animal models, so they can explore many more hypotheses at the same time.
“What we'd like to do in principle is to look at the entire set of candidate hypotheses at once, and then let the data tell us what hypotheses are compatible and not compatible with the data,” she said. “It’s a philosophical change in terms of how we can think about science that is only made possible because of the technological innovation.”
As she built on the idea, the SUNY Stony Brook Biomedical Engineering Professor, who also holds an affiliation with The Picower Institute, assembled a “dream team” of scientists and engineers including Miller, MIT Math Professor Alan Edelman, MIT Research Affiliate Chris Rauckaukas, Stony Brook Associate Professor Helmut Strey, and Dartmouth College Professor Rick Granger. Meanwhile, to launch the effort, Mujica-Parodi received funding from philanthropists David Baszucki and Jan Ellison Baszucki.
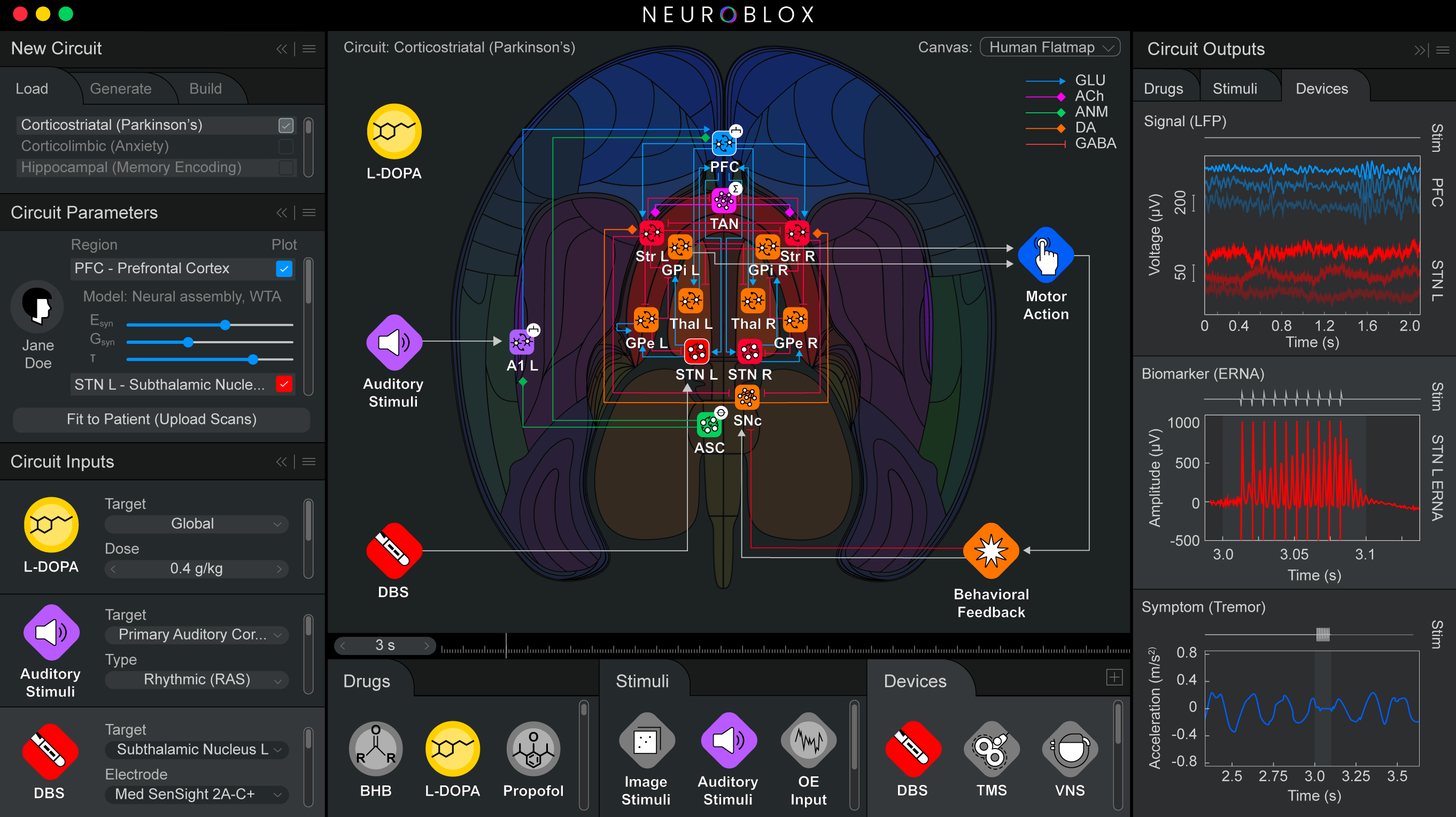
Neuroblox owes its name, in part, to how it’s built. Like an integrated circuit, it integrates fundamental computational components, also called “primitives” or “blox.” Unlike a computer chip, Neuroblox is biomimetic, meaning that the blox model fundamental processing units made up of spiking neurons, experimentally derived from neuroanatomical and neurophysiological data. These primitives are the elements from which larger-scale circuits are composed, which in turn are the elements from which even larger-scale circuits are composed.
This modular, nested approach—validated against experimental brain data at every scale—extends all the way up to whole-brain computation, Mujica-Parodi said. The virtual neurons in each primitive can respond to electrical or molecular inputs, such as external sensory stimuli, signals from fellow neurons, or neuromodulators such as serotonin, metabolites, or drugs.
The architecture enables the model to connect cellular and molecular activity all the way to cognition and behavior. For instance, in Nature Communications in December 2025, the team published the results of a critical test: They asked Neuroblox to learn a simple visual categorization task that Miller had once asked lab animals to learn. The model did so with almost exactly the same erratic progress that the animals exhibited. Underlying neural activity associated with this learning also closely resembled that of the animals. In fact, the model replicated and therefore revealed something that had remained hidden in the original lab animal data. Some cells, even as learning progresses, continue to inject error. Why? Miller speculates they might give the brain the flexibility to discover a new path forward if the rules of the task ever change.
While the study demonstrated sensory and neuromodulatory input, advances since then have enabled Neuroblox to accommodate more, Mujica-Parodi said. This has enabled the company to model anesthesia, deep brain stimulation, and even questions related to interactions between the central nervous system and other physiological systems, such as how dietary ketosis and diabetic insulin resistance affect the brain.
The model’s versatility makes it ideal for accelerating the discovery of therapeutics, Mujica-Parodi and Miller said, because it can reduce the guesswork that holds back many drug studies. For instance, while scientists have become good at crafting molecules to bind to target receptors on neurons, it has been slow going to figure out what will happen in brain circuits or to behavior once the binding occurs. By enabling rapid, scalable computational testing of hypotheses, Neuroblox can help answer such questions to improve drug trials, Mujica-Parodi said.
Many methods move neuroscience forward. New simulations, both on the lab benchtop and the computer desktop, promise to move it forward faster.