July 1, 2025
About Us
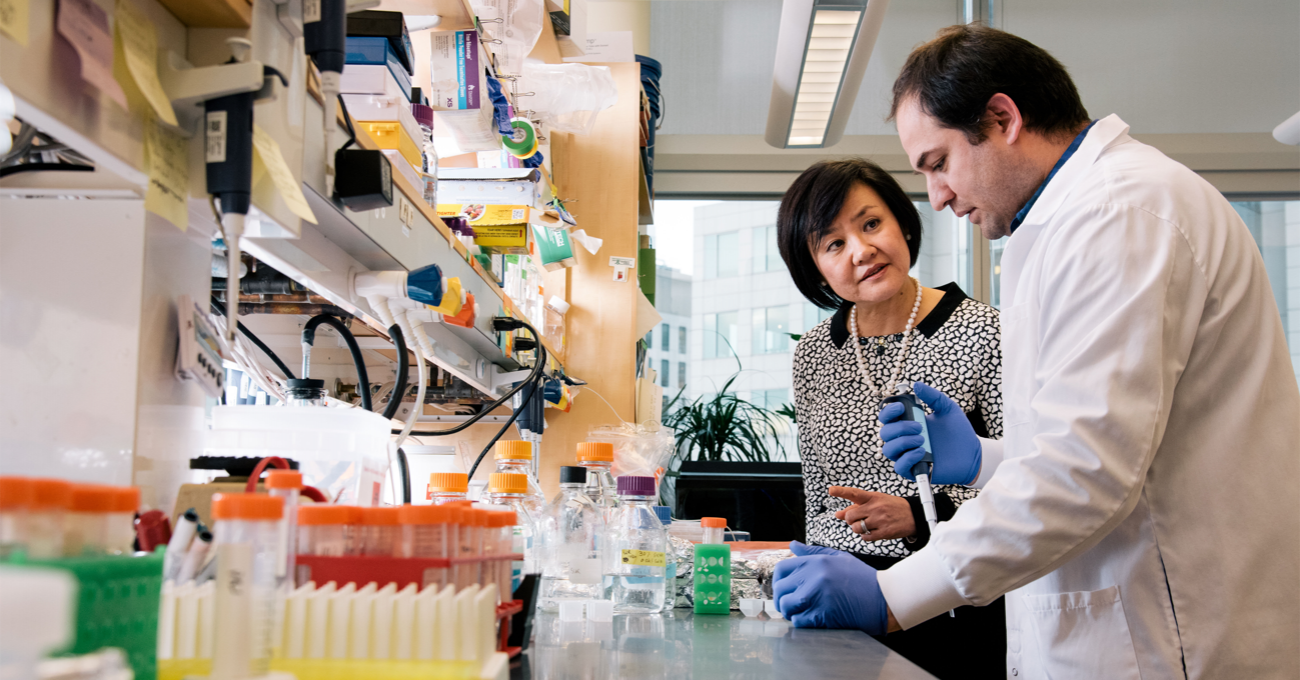
The Picower Institute is a community of scientists dedicated to understanding the mechanisms that drive learning and memory and related functions such as cognition, emotion, perception, behavior, and consciousness. Institute neuroscientists explore the brain and nervous system at multiple scales, from genes and molecules, to cells and synapses, to circuits and systems, producing novel insights into how disruptions in these mechanisms can lead to developmental, psychiatric or neurodegenerative disease.

The tremendous need for research to address the burdens of the aging brain—memory loss, cognitive decline, and dementia—is what gave rise to an Institute-wide call to action at MIT, the Aging Brain Initiative.